The importance of pedigree analysis in breeding
The Inbreeding Coefficient (IC) or Coefficient of Inbreeding (COI) or Coefficient of Kinship is the relationship of the parents of the individual. It is defined as the probability that the two genes this individual has in a locus are identical by descent, i.e., they are both inherited from a common ancestor. The calculation of the inbreeding coefficient is done up to a given pedigree depth (number of generations): the greater the depth of the pedigree, the greater the chance of finding new common ancestors, and the higher the inbreeding coefficient.
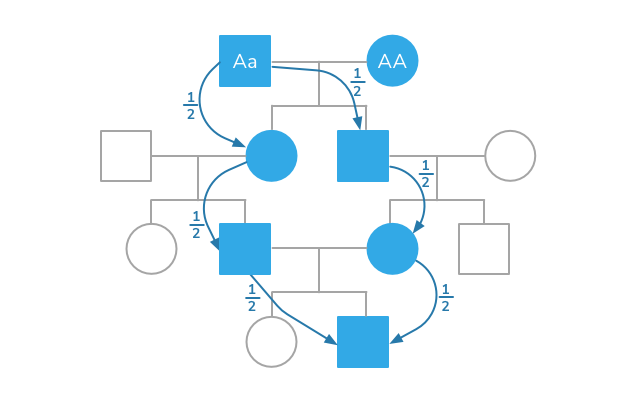
The Breed Archive allows you to choose from one to ten generations. We have chosen seven generations as the default setting — on the one hand, this is accurate enough and, on the other, fast enough, and there is a good chance that a large percentage of the pedigrees will be complete for that number of generations.
A dog receives a high COI if the same ancestors appear on both the sire and dam side. You can have a low COI for an offspring even in cases where both parents have high COIs - if sire and dam do not share the same ancestors, the offspring won't have a high COI. Therefore it is also important to have a look at the ancestor loss to get an idea of the genetic diversity of a particular dog or a planned litter.
Comparing COI
It makes sense to compare the inbreeding coefficients of different pedigrees only if the number of generations used for the calculation is the same and the pedigrees are complete for the generations. So please keep in mind that when you compare the COI of two or more potential matings (e.g. for future breeding), you need to do this with the same number of complete generations for all your test matings. The COI might increase substantially with the number of generations.
COI calculation
There are different methods to compute the COI; e.g. the path-coefficient method and the tabular method. While the path-coefficient approach is considered more complete and accurate, the tabular method is the more modern and faster algorithm for calculating inbreeding coefficients. For the Breed Archives we are using the tabular method to calculate the COI, based on the data available for a particular dog. As mentioned earlier, you should only compare COIs of different dogs that were calculated with the same method and with the same number of full generations.
Partial COI
The partial inbreeding coefficients, Fij, measure the probability that the alleles at an arbitrary locus in individual i are identical by descent and that the alleles were derived from an allele in founder j. The partial inbreeding coefficients of the ancestors of an individual can be seen as their "contribution" to this individual’s COI, i.e., in a full pedigree the sum of all partial COIs of all ancestors of the individual must equal the individual’s COI!
Ancestors contributing the most to an individual’s COI do not automatically have the highest inbred percentage. By being a closer relative (grandparent, great-grandparent) an animal with a low inbred percentage might have a larger contributing factor than other animals with a higher inbred percentage. The animal you selected for pedigree analysis will have most in common with the ancestors having the highest partial inbreeding coefficient.
Ancestor loss
The ancestor loss is calculated by comparing the number of possible ancestors in a pedigree with the actual number of different ancestors. For example, a four-generation pedigree contains 30 animals and if a particular pedigree consists of 30 unique animals then this pedigree has no ancestor loss at all; but in a pedigree with some inbreeding there might only be 26 unique animals in these four generations, meaning an ancestor loss of 13.3%.
It is important to note that if the dog’s pedigree is incomplete you won’t get reliable numbers for either the COI or the ancestor loss for generations higher than the last complete generation!
Blood quota
Blood quota is the percentage of a pedigree that is made up by any one ancestor. Sire and dam, obviously, each make up 50% of a pedigree; grandsires and granddams 25%; great-grandparents contribute 12.5%, etc.
E.g. an animal that appears in the grandparent generation contributes with 25%, if it also appears in the great-grandparent generation (12.5%) then it has a blood quota of 37.5%(25 + 12.5).
Why pedigree analysis is important
The pedigree analysis is most important for breeders when they are planning a mating of two individuals, as it provides information about the degree of relatedness of the ancestors of the pedigree, the expected genetic diversity and how much influence a single animal in the pedigree will have on the planned litter. The pedigree analysis also helps future owners to verify if their chosen puppy has enough genetic variability and if the breeder is avoiding the reduction of genetic diversity in the breed. (Low genetic diversity predisposes dogs to recessive hereditary diseases and may increase the risk of autoimmune diseases.)
On each dog’s entry in the Breed Archive you will find a link to its pedigree analysis page. The pedigree analysis currently provides information about the coefficient of inbreeding (COI), the ancestor loss and the list of ancestors with their blood quota. The partial COI calculation is available only up to the 5th generation.
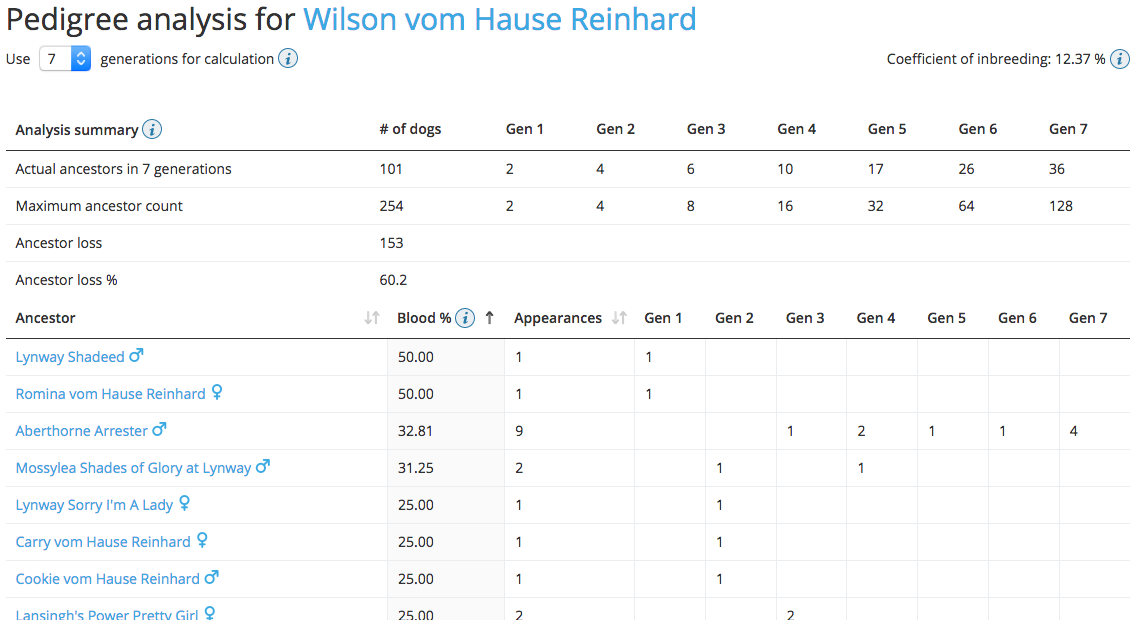
Pedigree analysis of Collie Wilson vom Hause Reinhard
Limitations of pedigree analysis for measuring genetic diversity
It is important to mention that measures like the coefficient of inbreeding or the ancestor loss are not the only ways to measure genetic diversity. Analysis of mitochondrial and microsatellite DNA are becoming increasingly popular. While measures like the COI are mathematical formulas, calculating probabilities that do not necessarily apply 100% for a particular dog (all litter mates have the same COI), the DNA analysis reveals actual diversity, i.e., with DNA analysis genetic diversity of litter siblings can vary.